NGS Library Quantification
准确量化载入流动槽中的可扩增文库分子的数量是二代测序步骤中获得高质量读段数据中关键的步骤。文库DNA载量不足会导致簇团密度低,测序产量降低。 文库DNA过多可能会增加簇团密度,导致数据质量差。 二代测序文库定量的标准方法,通过电泳或分光光度法,灵敏度低,对适配体结合的DNA无特异性,通常需要大量的文库样本进行分析。
qPCR具有较高的灵敏度和较宽的动态范围,因此被认为是二代测序文库定量的金标准,即使是对非常稀的文库也能准确地进行定量。
Library Quantification Kit Performance
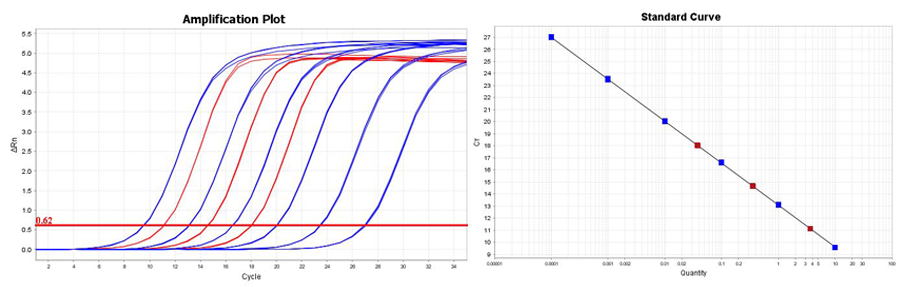
qPCR amplification of each of the six supplied DNA standards (blue) and a tenfold serial dilution of the diluted Illumina NGS library (red) were carried out with the supplied primer sets in triplicate. From the resulting standard curve, a size adjusted library concentration was then determined. The amplification plots demonstrate a limit of detection of 0.0001 pM (100 aM).
Description
Accurate quantification of the number of amplifiable library molecules loading into a flow cell is one of the most critical steps in the next-generation sequencing (NGS) workflow in obtaining high-quality read data with NGS technologies. Loading an insufficient amount of library DNA will result in low cluster density and reduced sequencing yield. An overabundance of library DNA may increase cluster density and result in poor quality data. Standard methods of NGS library quantification, by electrophoresis or spectrophotometry, have low sensitivity, are non-specific for adapter-bound DNA and typically require a large amount of library sample for analysis.
With its greater sensitivity and broad dynamic range, qPCR is therefore seen as the gold standard for NGS library quantification, as it accurately measures the number of molecules that can serve as templates during library and cluster amplification, even with very dilute libraries.
Specifications
| Description | Fast and robust quantification of Illumina-based NGS libraries. It consists of a SYBR®-based Fast qPCR Mix, pre-diluted DNA standards, primer mix and dilution buffer, designed to accurately measure the number of amplifiable DNA fragments before loading into a flow cell. |
| Concentration | 2x |
| Format | Clear, colorless solution |
| Hot Start | Antibody mediated |
| Application | qPCR, two-step RT-qPCR |
| Sample type | cDNA, DNA |
| Presentation | 9 vials |
| Storage | -20 °C |
| Mix stability | See outer label |
| Consistency | ± 0.5 Ct variance between test and reference sample |
| DNA Contamination | None detected in PCR amplification with traces overlay with the negative control on E. coli and mouse genomic DNA specific targets. |
| DNase Contamination | No detectable degradation |
产品资料
无甘油 T4 DNA 连接酶无甘油 T4 DNA 连接酶
High-Specificity Pfu HS CatalogHigh-Specificity Pfu HS Catalog
FAQs: NGS Library Quantification
With its greater sensitivity and broad dynamic range, qPCR is seen as the gold standard for NGS library quantification, as it accurately measures the number of molecules that can serve as templates during library and cluster amplification, even with very dilute libraries.
准确量化载入流动槽中的可扩增文库分子的数量是二代测序步骤中获得高质量读段数据中关键的步骤。文库DNA载量不足会导致簇团密度低,测序产量降低。 文库DNA过多可能会增加簇团密度,导致数据质量差。 二代测序文库定量的标准方法,通过电泳或分光光度法,灵敏度低,对适配体结合的DNA无特异性,通常需要大量的文库样本进行分析。 qPCR具有较高的灵敏度和较宽的动态范围,因此被认为是二代测序文库定量的金标准,即使是对非常稀的文库也能准确地进行定量。
No. However, the Library Quantification Kit contains primers which are compatible with P5 and P7 adaptor sequences specific to the Illumina platform.
Yes, the prepared library has to be diluted prior to running qPCR. Generally, we recommend preparing a 1:10,000 1:100,000 and 1:1,000,000 diluted sample of the library using the Dilution Buffer provided. Using two or more different dilutions of the library will ensure that at least one dilution falls within the dynamic range of the standard curve generated. This could be especially useful when the library concentration is high.
Instruments like the Qubit, spectrophotometer and Bioanalyser measure the total amount of DNA present in solution. The Library Quantification Kit will only quantify molecules with both adapters present. If the concentration measured by qPCR is significantly lower than what is measured by another methods, this suggests that only a small proportion of molecules correctly incorporated the adapters during adapter ligation.
Yes, we always recommend running triplicate qPCRs, as qPCR is an extremely sensitive measurement technique that is vulnerable to variation arising from liquid handling, instrument performing and sampling error.
Yes. The intercalating dye used in the kit binds DNA in proportion to the number of base pairs per double-stranded DNA product, so it is necessary to correct for product length by normalizing to the size of the DNA standard, 342 bp. The formular for doing this correction can be found in the Product Handling Guide.
与我们的专业团队联系
想了解更多迈迪安免疫和分子产品信息?欢迎与我们联系