Lyo-Ready™ qPCR and RT-qPCR Master Mixes
Meridians Lyo-Ready™ master mixes are glycerol-free mixes, ideal for developing ambient temperature, lyophilized, multiplex qPCR and RT-qPCR molecular diagnostic tests and are suited for high-throughput, automated platforms or microfluidic qPCR platforms requiring millisecond rehydration times.
Have questions about a product?
Contact us to learn more about Meridian’s molecular or immunoassay reagent portfolio. We want to hear from you!
High efficiency and sensitivity from both DNA (red) and RNA (blue) templates
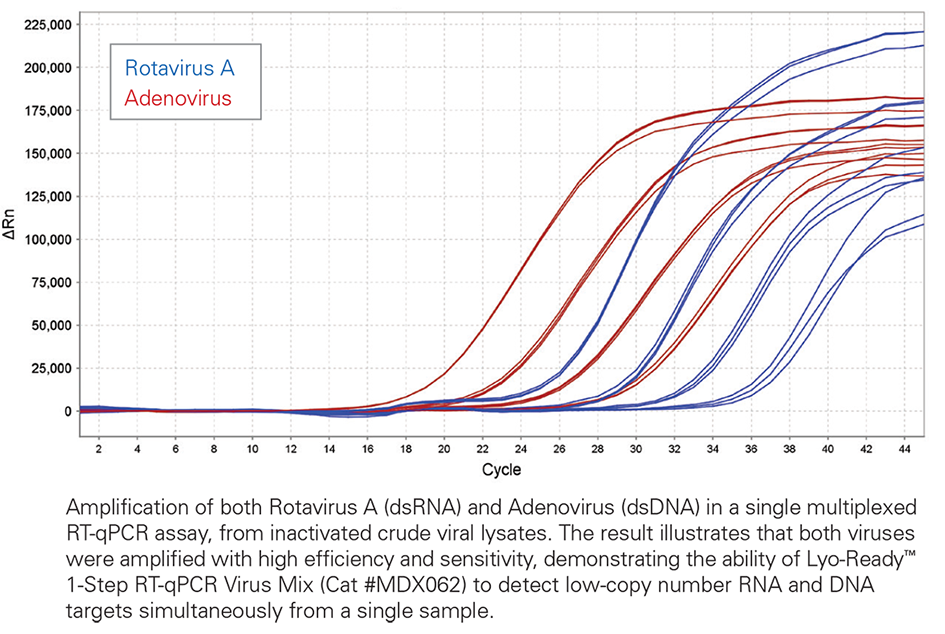
Lyo-Ready qPCR Mix, MDX021
Available in 5 mL (500 Rxns) or 100 mL (10,000 Rxns) aliquots
Lyo-Ready qPCR Mix 2.6x, MDX023
Available in 8 mL (500 Rxns) or 100 mL (12,500 Rxns) aliquots
Lyo-Ready 1-Step RT-qPCR, MDX024
Available in 10 mL (1,000 Rxns) or 100 mL (10,000 Rxns) aliquots
Lyo-Ready 1-Step RT-qPCR Virus, MDX062
Available in 10 mL (1,000 Rxns) or 100 mL (10,000 Rxns) aliquots
Molecular diagnostic tests are progressively moving towards lyophilized formats. There are several advantages for this, including ambient temperature shipping and storage, extended shelf-life, reduced operating steps and potential errors and increased flexibility in sample volume. In order to be compatible with drying, however, enzyme preparations must be glycerol-free and include specialized excipients that preserve the mixture as it is exposed to various conditions including freezing, vacuum, and dehydration.
Lyo-Ready™ qPCR Mix (2x) and Lyo-Ready™ qPCR Mix 2.6x Mix are used for the development of DNA based tests multiplex qPCR, enabling highly sensitive detection of target DNA, the Lyo-Ready™ qPCR Mix 2.6x Mix is at a slightly higher concentration for use with microfluidic qPCR, where the initial volume needs to be smaller.
Lyo-Ready™ 1-Step RT-qPCR and Lyo-Ready™ 1-Step RT-qPCR Virus Mix are 2x RT-qPCR master mixes and can be used for the development of both RNA and DNA based molecular diagnostic tests, they contain a separate lyo-compatible MMLV-RT and enzyme dilution buffer. The Lyo-Ready™ 1-Step RT-qPCR Virus Mix reverse transcriptase is also more thermostable and has been optimized for amplification of RNA or DNA viruses with a high secondary structure (reverse transcriptase remains active at 55-60 °C).
Catalogs & Brochures
Lyo-Ready 1-Step RT-qPCR MixesLyo-Ready 1-Step RT-qPCR Mixes
Master Mixes for Molecular Ambient-Temperature Stable AssaysMaster Mixes for Molecular Ambient-Temperature Stable Assays
FAQs: Lyo-Ready qPCR and RT-qPCR Master Mixes
Slope of standard 10-fold dilution curve indicates qPCR efficiency. The efficiency of the qPCR should be between 90–110% (−3.58 ≥ slope ≥ −3.1). If the efficiency is greater than 100%, this indicated inaccurate sample and reagent pipetting, if it is less than 100%, this indicates your samples may contain PCR inhibitors or your qPCR primer and/or probe design may not be optimal.
One-step reaction
• Accurate representation of target copy number
• Simple and rapid
• Fewer pipetting steps (reducing possible errors and contamination)
• Best option for high-throughput screening
• Best method when only a few assays are run repeatedly
• Multiplex qPCR of gene of interest and control can be done in single well, from same RNA sample
Two-step reaction
• Two buffers optimized for independent RT and qPCR
• Highly sensitive
• Potentially more efficient because random primers and oligo d(T) can be used
• Possibility to stock cDNA to quantify several targets
• Recommended when the reaction is performed with a limiting amount of starting material
So one-step workflows are commonly favoured in molecular diagnostic tests. Two-step RT-qPCR is preferred when multiple interrogations will be made of the same starting material or where archiving of cDNA may be required.
Yes, reaction conditions are the same and they will produce the same Ct values, even with multiplex qPCR assays. This means that the liquid mixes can be used to create SOPs if you do not want to lyophilize and then later if lyophilization is required (for example to increase sensitivity), the SOP can be updated for lyophilization, it does not require completely new SOPs to be written.
Many fluorescent qPCR primer- and probe-based chemistries have been devised and are available from different commercial vendors for molecular diagnostic tests, the two most commonly used are:
• Hydrolysis (TaqMan) probes. The main advantages of using hydrolysis probes are high specificity, a high signal-to-noise ratio, and the ability to perform multiplex qPCR reactions. The disadvantages are that the initial cost of the probe.
• Molecular beacons. Molecular beacons are highly specific, can be used for multiplexing, and if the target sequence does not match the beacon sequence exactly, hybridization and fluorescence will not occur. Unlike hydrolysis probes, molecular beacons are displaced but not destroyed during amplification and can be used for melt curve analysis if necessary. The main disadvantage is that they are difficult to design, requiring a stable hairpin stem that is strong enough that the molecule will not spontaneously fold into non-hairpin conformations but not be too strong, or it may not properly hybridize to the target.
The Lyo-Ready™ qPCR and RT-qPCR master mixes have some inhibitor tolerance, but for amplification from crude lysates or inhibitor-rich samples we have the universal Inhibitor-Tolerant qPCR/RT-qPCR mixes and for even greater sensitivity, we have the sample specific (blood, saliva, urine, stool, and plant) mixes.
No-RT control reactions are useful for determining issues that may arise from the amplification of genomic DNA that may be present in a sample. However, many RNA transcripts are present at low abundance and for these transcripts, it is recommended to perform a No-RT control reaction if primer sets do not span exon-exon junctions. If the primers do span exon-exon junctions, or genomic DNA is not likely to interfere (when looking at RNA viruses in a sample for example) a No-RT control is not required.
Following lyophilization in the presence or absence of primers and probes, the Lyo‑Ready™ qPCR and RT-qPCR master mixes are stable for a minimum of 24 months at ambient temperature (17 – 23 °C). Following rehydration, the assay reproducibility, sensitivity and robustness will be the same as for a freshly made liquid mix, making the mixes ideal for point-of-care molecular diagnostic tests and microfluidic qPCR devices.
Get In Touch With A Specialist
Have questions about a product? Want to learn more about Meridian’s molecular or immunoassay reagent portfolio? We want to hear from you!