Home » Life Science » Products » Molecular Reagents » DNA-RNA Extraction Controls » RT-qPCR Extraction Control Red
RT-qPCR Extraction Control Red
Molecular IVD companies continue to experience tighter and tighter regulations in regard to incorporating appropriate positive and negative controls into their assays. Meridian’s RT-qPCR Extraction Control Red uses artificial micelles of a known concentration that contain quantitative PCR positive control RNA sequence (with no known homology to any organism, so that it does not interfere with the detection of the sample target RNA). Their composition closely resembles a test sample and serves as a full-process quantitative PCR positive control, monitoring the success of an RT-qPCR assay from lysis/extraction to reverse transcription and amplification
Have questions about a product?
Meridian's RT-qPCR Extraction Control Red
- Extraction control undergoes the same processing as test sample
- Confirms successes of extraction step and monitors co-purification of PCR inhibitors
- Probe based designed specifically for multiplex quantitative PCR assays
- Quantitative PCR positive control sequence is unique with no homology to any organism
Workflow
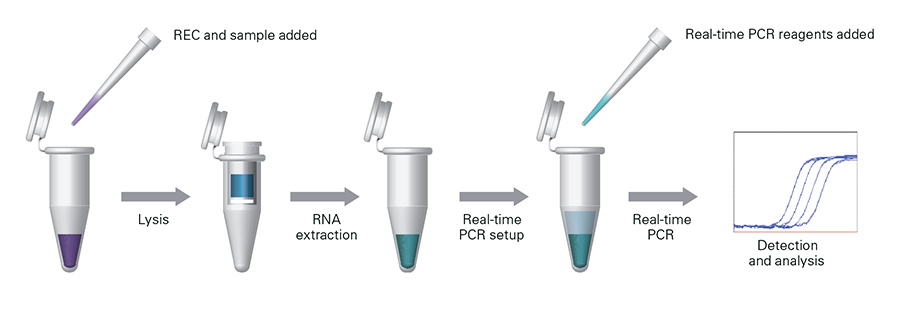
RT-qPCR Extraction Control Red, MDX028
Control used to monitor the success of an RT-qPCR assay from lysis/extraction to amplification and reduces the chance of obtaining false negative results from the sample RNA.
Documents & Resources
Description
RT-qPCR Extraction Control Red contains an internal control RNA sequence, with no known homology to any organism, in an artificial cell. The RT-qPCR Extraction Control is spiked in the with lysis buffer into the sample prior to RNA extraction. Following RNA extraction, the control mix (containing specific primers and probe (emission wavelength = 670nm)) is added to the extracted RNA prior to amplification. The detection of RT-qPCR Extraction Control acts as a PCR positive control and confirms the success of the extraction, reverse transcription, and amplification steps, avoiding the misinterpretation of a false negative result.
Specifications
| Description | Control used to monitor the success of an RT-qPCR assay from lysis/extraction to amplification and reduces the chance of obtaining false negative results from the sample RNA. |
| Concentration | 25x |
| Appearance | Clear, colorless solution |
| Internal Control RNA | In micells |
| Control Mix | Primers and Quasar® 670 labeled probe |
| Application | Probe-based, one-step RT-qPCR |
| Sample type | Tissue, cells |
| Presentation | 3 vials |
| Storage | -80 °C |
| Mix stability | See outer label |
| Functionaity/ Consistency | Ct of 27 – 30. The difference between test and history is less than 1 Ct. |
Catalogs & Brochures
Related Products
FAQs: RT-qPCR Extraction Control
What are the differences / advantages in comparison with other methods like measurement of housekeeping gene, internal control / spike in controls?
The benefit of the artificial RT-qPCR Extraction Control in evaluation of the extraction process is the possibility to validate sample extraction process. Signal derived from the RT-qPCR Extraction Control confirms the success of the extraction step and monitors co-purification of PCR inhibitors that may cause biased or PCR false negative results. Housekeeping genes or spike in controls do not monitor the whole extraction process, especially the lysis step and in the case of housekeeping genes, extraction efficiency and degraded samples as well as the possibility of the housekeeping gene expression level changing.
What is internal RNA control in RNA extraction?
When performing RNA extraction, it is often advantageous to have an exogenous source of RNA template that is spiked into the sample prior to lysis. This control RNA is then co-purified with the sample RNA and can be detected as a PCR positive control for the whole extraction process.
Do I have to have an internal PCR control?
Although it is not mandatory yet, regulators across the world are tightening up recommendations for the addition of internal controls, EU: IVDD to IVDR 2017/746 “Control materials with quantitative or qualitative assigned values intended for one specific analyte or multiple analytes shall be classified in the same class as the device.” MDSAP with FDA (“general controls ensuring effectiveness of the device”), TGA (“provisions for the user, at time of use, of device performance”), MHLW and ANVISA
How careful do I need to be with the RT-qPCR Extraction Control?
The RT-qPCR Extraction Controls are in micelles and so care needs to be taken when handling them. We recommend storing the RT-qPCR Extraction Control at -80 ᵒC and adding it to the lysis buffer before adding to the sample.
Do I need to supply the primers and probe for the RT-qPCR RNA Extraction Control?
The primers and probe for the RT-qPCR Extraction Controls are in the Control Mix.
Why are there two colors for the RT-qPCR Extraction Controls?
The two colors are for the fluorescent dyes on the RT-qPCR Extraction Control probe. Control Mix Red (Cy5 – emission wavelength = 670nm) and Control Mix Orange (HEX – emission wavelength = 560nm). This allows you to chose one that will fit with your existing protocol in a multiplex RT-qPCR assay.
Do I need a specific extraction method for the RT-qPCR Extraction Control?
No, the RT-qPCR Extraction Control will work with commercially available silica-membrane extraction kits and CHELEX matrices and has been tested on a wide range of qPCR platforms.
Get In Touch With A Specialist
Have questions about a product? Want to learn more about Meridian’s molecular or immunoassay reagent portfolio? We want to hear from you!
By submitting your information in this form, you agree that your personal information may be stored and processed in any country where we have facilities or service providers, and by using our “Contact Us” page you agree to the transfer of information to countries outside of your country of residence, including to the United States, which may provide for different data protection rules than in your country. The information you submit will be governed by our Privacy Statement.